DR. YOUSIF SHAMOO
Shamoo Laboratory
Antimicrobial resistance is a global crisis.
There is an urgent need for new antibiotics as well as strategies to combat resistance, particularly in hospital settings where drugs of last resort like daptomycin (DAP) are becoming ineffective. A critical step in the development of such strategies lies in understanding the evolutionary trajectories responsible for resistance and the identification of proteins or biochemical pathways that can be good targets for the design of novel treatment strategies and co-drugs. I am interested in the underlying physicochemical principles of adaptation within bacterial populations during evolution. We use a combination of experimental evolution and biophysical approaches to link changes in protein structure and function to their resulting phenotypes within evolving populations (genotype to phenotype to physicochemical mapping). We go on to extend these physicochemical principles to predict the success or failure of specific adaptive alleles undergoing selection. By combining approaches from biophysics, genomics, and experimental evolution, identify and characterize successful evolutionary trajectories and then link those intermediates to the evolutionary trajectories of bacterial populations. More recently, we have combined our experience with quantitative experimental evolution and biophysics to develop new microfluidic approaches to elucidate the biochemical basis of adaptation to antimicrobials and for drug discovery. New Projects! More recently, we have begun to use microfluidics and approaches from synthetic biology to explore the vast chemical diversity of Streptomyces strains for antimicrobial drug discovery and for applications in biosurveillance.
|
Daptomycin resistance in the clinically importantl pathogen Enterococci is mediated by the LiaFSR signaling pathway. LiaFSR is a cell envelope stress response pathway. Together with the Arias group at Methodist Research Institute, we have studied the acquisition of Daptomycin resistance. Over many years of hard work (that includes other groups), the multilayered defense of Enterococci against Daptomycin and other membrane stress agents has been mapped out.

|
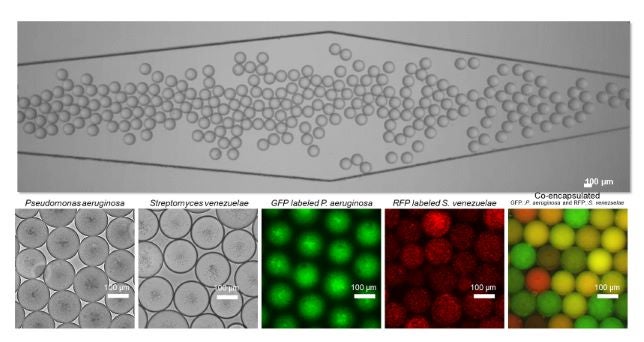
Other organisms of interest to the lab include E. faecium, Acinetobacter baumannii, Pseudomonas aeruginosa, Nocardia and Francisella. In addition to determining the molecular mechanisms leading to resistance, the lab is also interested in studying microbial interactions. With the help of microfluidic techniques to create spatially segregated emulsion systems, we wish to understand bacterial interactions and the evolutionary processes that lead to such relations. For more information and details of the projects going on in the Shamoo lab, please view the “Current lab members” and Shamoo Lab publications pages. |
Shamoo Lab News
3/27/2026
Aaron Pergler wins Barry Goldwater Scholarship!
11/14/2025
Aaron Pergler wins Excellence in STEM Communication Best Presentation Award at San Jacinto College Research Symposium!
10/29/2024
Aaron Stibelman wins McSherry-Poe research award!
09/01/2024
Kat Rhatigan wins "Top Presenter" at the Rice University Research Symposium! Congratulations Kat!
04/29/2024
Dr. Shamoo elected American Association for the Advancement of Science (AAAS) Fellow! link
10-17-2023
New publication!! Congratulations to Saoirse on her new publication using microfluidics to study the evolutionary trajectories of Pseudomonas aeruginosa in microaerobic environments. Link.
9-8-2023
New publication!! Congratulations to Dr. Xinhao Song on his new publication, which introduces a non-optical detection method utilizing methyl halide gases for the quantification of filamentous bacteria in microdroplets! Link.
4-4-2023
Congratulations to graduate student Adeline Supandy for successfully defending her PhD thesis!
3-20-2023
Congratulations to graduate student Xinhao Song for successfully defending his PhD thesis!
03-17-2022
New publication!! Congratulations to Dr. Ramya Ganiga Prabhakar for her paper in establishing parameters for enrichment of rare bacterial subpopulations in microdroplets! Link.
1-20-2022
New publication!! Congratulations to Dr. Heer Mehta for her paper in investigating Francisella tularensis subsp. holarctica antibiotic resistance in macrophages! Link.
1-14-2023
New publication!! Congratulations to graduate student Yue Zhou for her paper in characterizing Enolpyruvate transferase MurAAA149E Link.
November 2022
Congratulations to Dr. Heer Mehta to her new position at Cemvita!
8-8-2022
IBB Best Poster Award Congratulations to Lauren LeDay for winning Best Poster at the IBB summer research event! Link.
5-11-2022
New publication!! Congratulations to graduate student Adeline Supandy for her paper identifying the mechanisms of resistance in Enterococcus faecium to daptomycin and fosfomycin Link.
5-9-2022
New publication in collaboration with Craig Miller and Chris Marx from UIdaho! Mutational Switch-Backs Can Accelerate Evolution of Francisella to a Combination of Ciprofloxacin and Doxycycline. Link
4-21-2022
New publication!! Congratulations to Post-doc Seokju Seo and the entire microfluidics team for the new Star Protocols paper.
3-29-2022
12-28-2021
New paper out! Congratulations to everyone involved in establishing the microfluidics platform for conducting experimental evolution in our lab. Check out the first of many papers on this topic here.
11-16-2021
Congratulations to Ramya for a successful thesis defense!
5-26-2021
New publication in collaboration with Chris Marx at University of Idaho! EfgA is a conserved formaldehyde sensor that leads to bacterial growth arrest in response to elevated formaldehyde
May 2021
Dr. Yousif Shamoo in his own words https://magazine.rice.edu/2021/05/the-experimentalist/
February 2021
Post-doc position open in Shamoo Lab https://jobs.rice.edu/postings/25590
01-19-2021
New publication: Daptomycin resistance in Enterococcus faecium can be delayed by disruption of the LiaFSR stress response pathway. DOI: 10.1128/AAC.01317-20
10-12-2020
Check out Dr. Shamoo's podcast on "Hold These Truths with Dan Crenshaw" about Innovating the Future: The Next Frontiers In Clean Energy, Vaccines and Agriculture!!
3-5-2020
Check out our latest PLoS Pathogens review article on pathogenic Nocardia - https://doi.org/10.1371/journal.ppat.1008280
8-12-19
Shamoo Lab on EurekAlert!
https://www.eurekalert.org/pub_releases/2019-08/ru-rus081219.php?fbclid=IwAR0isffII7jMz5hmdmGCezWXQzdkUkJwjz_a577bRp8Vl1zciCg_6ESK4Z4
7-17-19
New publication: Environment Shapes the Accessible Daptomycin Resistance Mechanisms in Enterococcus faecium. DOI: 10.1128/AAC.00790-19
7-15-19
Congratulations to graduate student Amy Prater for successfully defending her PhD thesis!
June 2019
Congratulations to postdoc Zaigao Tan for getting a faculty position at Shanghai Jiao Tong University